Description
Reminders:
1. Out of 100 points. 5 Questions. Contains 8 pages.
2. If you use Late days, mark how many you are using (out of maximum 4 available) at the top ofyour answer PDF.
3. There could be more than one correct answer. We shall accept them all.
4. Whenever you are making an assumption, please state it clearly.
5. You will submit a solution pdf LASTNAME.pdf containing your answers and the plots as well as a zip file LASTNAME.zip that contains your code and any output files.
7. Additionally, you will submit one zip file LASTNAME.zip that contains your code and any results files. Code and results for each question should be contained in a separate sub-directory (Eg: Q1) and there should be a README.txt file for each sub-directory explaining any packages to install, command to run the code files and location of the expected output. Please follow the naming convention strictly.
8. If a question asks you to submit code please enter the file path (Eg: Q1/Q-1.3.1.py) in the solution pdf.
9. If needed, you can download all the datasets needed for this homework from ICON. Informationabout the datasets will be given in the README.txt file.
1. (50 points) SIR Model
We observe that at the end of a very long semester (t → ∞) 60% of the student population are interested in CS:4980. What is the expected fraction of fellow students an interested (infected) student spreads their interest to?
Note: Assume the initial infected 2% are all still contagious. (i.e., R(0) = 0)
Solution:
Q 1.2 (13 points) ODE ⇔ Networks Let us try to derive the SIR ODE model from the SIR network model. Assume that the network is a clique of size N (i.e. everyone is connected to everyone else with a total population of N individuals). Further assume the model evolves in discrete time steps. At each step:
1. each susceptible individual u picks a person v from its neighbors, uniformly randomly
2. if v is infected, then u gets infected with probability β.
3. each infected individual goes into the R state with probability γ
Note: This SIR network model is slightly different from that described in class which was from the point of view of infected node. Here, we describe from the point of view of a susceptible node but produces equivalent result.
Let X(t) be the number of individuals who are in the S state at a time t. Similarly, Y (t) is the number of individuals who are in state I and Z(t) be the number of individuals in state R.
Q 1.2.1 (2 points) What is the probability P(t) of a single susceptible individual getting infected at time t? Clearly the total number of expected new infections at time t + 1 will be P(t) × X(t).
Solution:
Q 1.2.2 (4 points) What is the number of expected susceptible individuals at time t + 1 i.e. what is E[X(t + 1)]? Write it down first in terms of P(t) and X(t) and then substitute P(t) from Q 1.2.1. Similarly, write down E[Y (t + 1)] and E[Z(t + 1)].
Solution:
Q 1.2.3 (2 points) Assuming E[X(t)] ≈ X(t) , write down the expression for X(t+1)–X(t)? Similarly, write down Y (t + 1)–Y (t) and Z(t + 1)–Z(t).
Solution:
The steps above assumed 1 unit of time. Let us change the unit of time to a small ∆t instead. Hence, β should now become β∆t (ditto for γ).
Q 1.2.4 (2 points) Write down X(t+∆t)–X(t),Y (t+∆t)–Y (t), and Z(t+∆t)–Z(t), using your answers in Q 1.2.3 and the change mentioned above.
Solution:
Q 1.2.5 (3 points) Starting from your answers in Q 1.2.4, derive the standard SIR ODE equations you have seen in class. Make sure you show all the steps.
Solution:
Q 1.3 (32 points) Implementation and Calibration. In this question you will implement standard SIR ODE and SIR network models.
Q 1.3.1 (6 points) Implement the standard SIR ODE model (lecture 2, slide 19-24) in your favorite language [We strongly recommend python]. Your code should take in the β and γ values, and the initial fractions S(0),I(0),R(0), and max time as input, and give the fraction of infections, susceptible and recovered population (i.e. list of tuples (t,S(t),I(t),R(t)) for t = [0,…,max time]) at each time-step till max time. Submit the code.
Set the values of parameters as β = 0.1,γ = 0.05 and initiaize S(0) = 0.95,I(0) = 0.05,R(0) = 0. Show SIR curves (plot of S(t),I(t),R(t) over time) for upto T =
200.
Now set β = 0.05,γ = 0.1 and plot the SIR curves. Do you notice any difference in nature of the curves? Explain.
Solution:
Q 1.3.2 (10 points) Implement the discrete-time SIR Network model in your favorite language. Your code should take a graph-edge-list, the total number of nodes in the graph N, the β value, the γ value, the initial fractions S(0),I(0),R(0), and the max time as input, and give the fraction of infected nodes I(t), susceptible nodes S(t) and recovered nodes R(t) at each time-step till max time. Use the graph-edgelist from example.txt. You can set any random set of nodes to I/R/S states at time t=0 to reach the desired fraction. Submit the code and SIR plots. Also provide SIR plots for the network simulation with a complete graph of nodes N = 1000 given the initialization parameters from previous question Q 1.3.1. (As the SIR network model is stochastic, make sure you run the simulation 50 times, and take the average to get I(t),S(t),R(t). Further you can use packages to construct graphs like the networkx Python package or SimpleGraph, LightGraph Julia packages.)
Solution:
T
L = X(R(t) − Robserved(t))2
t=1
as given in slide 8, lecture 4 (assume the R state corresponds to mortality).
To initialize the boundary conditions, we set the initial values of I(0) as the fraction of cases and R(0) as fraction of deaths on the first day from the dataset. Set the initial value of S(0) as S(0) = 1–R(0)–I(0). Now calibrate the SIR model to find the best set of parameters β,γ that minimizes the objective function L.
Write down the values of β, γ you find after calibration. Plot S(t),I(t),R(t) vs time (just like the plots we saw in slide 19, lecture 3). Also plot another figure showing Robserved(t) you used to calibrate and the resultant R(t) after calibration. Submit the plots.
Solution:
Q 1.3.4 (8 points) Run the SIR Network model on a clique of size N = 100. Set the β and γ values as the ones you get in Q 1.3.3. Set the max time as 900. For the simulation, set the initial number of infected individuals I0 = ⌊I(0) ∗ N)⌋ (where I(0) is value you set in Q1.3.3), initial number of susceptibles S0 = ⌊(S(0) ∗ N)⌋, and the remaining nodes as R0 (i.e. R0 = N −⌊I(0) ∗ N⌋−⌊S(0) ∗ N⌋).
Note that ⌊x⌋ is the function which just rounds x to closest integer less than x. Plot the S(t),I(t),R(t) vs t on a plot as in Q1.3.3 (submit the plot). Repeat the same steps for N = 1000 and N = 10,000. Submit the plots. Compare the plots for different values of N and for the ODE (from Q 1.3.3). What do you observe?
Solution:
2. (5 points) Cayley Trees
Consider a symmetric k-regular tree in which each vertex is connected to the same number k of others, till we reach the leaves. This is also sometimes called a ‘Cayley’ Tree.
For example, if k = 3 the network will look like Figure 1.
Figure 1: k-regular Cayley tree for k = 3
Q 2.1 (2 points) In a k-Cayley tree, what is the number of nodes reachable in d steps from the central node? (e.g., the number of nodes reachable in d = 1 steps in the figure above is equal to 3). Show your derivation.
Note: A node u is reachable from the center node in d steps if there is a path of exactly d steps from center node to node u.
Solution:
Q 2.2 (3 points) What is the diameter of such a network (in terms of k and n)? The diameter of a graph is defined as the longest shortest path in the graph. Let the length of the shortest path between a pair of vertices be called the distance between the vertices. The maximum distance between any pair of vertices is called the diameter of the graph.
Please show your steps. Here n is the total number of nodes in the graph.
Solution:
3. (20 points) Calibrating IC Models
We will work with the following two datasets (download from canvas):
1. Network: network.txt Note that the graph is directed. Edge v → u is denoted as
(v,u).
2. Cascades/Action log: Ratings.timed.csv .
This file contains an action on each line. An action is a user rating a movie. That is, if a user v rates “The King’s Speech”, and then later v’s friend, say u, does the same, we consider the action of rating “The King’s Speech” as having propagated (cascaded) from v to u. In epidemiological terms you can think of v ‘infected’ u.
In this setting, the goal of the calibration procedure is to find out the probability pvu over each edge (v,u).
Q 3.1 (5 points) Let Av2u be the total number of actions of node v which cascaded to node u later. An action cascades from v to u if both the following conditions satisfy:
1. v must have taken the action before u.
2. Edge (v,u) exists. In other words, v is an in-neighbor of u.
Similarly, let Av be the total number of actions of node v in the database. Then you can think of node v making Av tries to infect node u, of which only Av2u tries succeeded. Write down the probability L of Av2u tries succeeding out of Av tries, assuming the probability of success is pvu. L is also called the likelihood of the data.
Solution:
We find the Maximum Likelihood Estimator . i.e
= argmax .
pvu
(Try to derive it on you own.)
Q 3.2 (15 points) Now we calibrate the IC model to estimate p∗vu for each edge (v,u). Using the two datasets given above, write code to calculate the for each edge (v,u) of the network. The code should take in the two datasets as input, and print “ “ (without quotes) on each line for each edge (v,u). Submit both the code and the output you get after running it on the two datasets.
Note 3: There are multiple ways to do this efficiently. We will not grade on efficiency of your code. However, it might be interesting for you to hear that you can do this in < 2 scans of the action/cascades log.
Solution:
4. (20 points) Zombie Apocalypse
There is a new disease outbreak in a town that turns people into zombies.
Q 4.1 (5 points) Stopping the outbreak using branching process. In the initial stage of the outbreak, the zombies don’t harm humans, they only infect them with the disease. The town has set aside 2 million dollars to deal with this outbreak which is currently in its infancy. The public health officials have two measures to slow down the outbreak:
1. Controlling how many people a zombie comes in contact with: If the officials spendx dollars, they expect a zombie to come in contact with 40 people.
2. Controlling the probability of transmission: If the officials spend y dollars, they expect the probability of a zombie transmitting the disease to another person it comes in contact with is 0. dollars.
What is the best way to allocate the 2 million dollar budget?
(Assume that most people are uninfected. Therefore, number of people an infected zombie comes in contact with is assumed to be constant.) Hint: Refer to Lecture 4.
Solution:
Q 4.2 (15 points) You failed to stop the outbreak. Now the zombie disease has turned sinister. The infected zombies don’t merely turn humans to zombies, instead they consume them.
Q 4.2.1 (6 points) Assume x(t) is the population that is human and y(t) the the population that are zombies at time t. We use the Lotka-Volterra Model (LV M) to model the dynamics of the populations:
(1)
(2)
Similar to Q 1.3.1, implement the ODE model above that takes in parameters α,β,γ,δ and initial states x(0),y(0) and returns fraction of populations x(t),y(t) till time-step max time. Submit the code for the model.
Set α = β = γ = δ = 1. Now set the values of initial population x(t) = 5,y(t) = 2 and plot the values of x(t),y(t) vs. t till T = 100. Submit the plot. Now experiment with two other initializations of x(0),y(0) for which the values of x and y don’t change over time.
Solution:
Q 4.2.2 (3 points) The zombie disease has mutated further! Now you realize that there are actually two variants of zombies that are consuming humans. Assume x(t) is the population that is human and y(t) the the population that are zombie variant 1 and z(t) the the population that are zombie variant 2 at time t.
We model the population dynamics as a variant of LV M called LV M2:
(3)
(4)
(5)
What are the fixed points of this LV M2 model?
Solution:
Q 4.2.3 (5 points) Implement the model in your favourite language that takes in parameters α,β,γ,δ,ϕ,ρ,ϵ and initial states x(0),y(0),z(0) and returns fraction of populations x(t),y(t),z(t) till time-step max time. Submit the code.
Assume α = β = ϕ = γ = ρ = 1, δ = 1.5 and ϵ = 2. For x(0) = 4,y(0) =
2,z(0) = 5 plot x(t),y(t),z(t) vs. t till T = 100. Also plot for other initializations of x(0),y(0),z(0) for which they are fixed points and show that the population doesn’t change over time. Submit the plots.
Solution:
Q 4.2.4 (1 point) Are the LV M2 plots periodic?
Solution:
5. (5 points) Closing Triangles
Imagine a social network consisting of the students in this class (if x knows/is friends with y, then there is an undirected edge between x and y). Let us create a subgraph centered around you. First, list down all the people from the class you know. These students are your neighboring nodes in the graph. Now, for each of your neighbors, ask them to list down the people they know from the class. Connect these people to your subgraph as well. These are your ’friends-of-friends’ (f-o-f).
Naturally some of your f-o-f will also be your friends. These create ’triangles’ centered around you in the graph.
1. Draw the constructed subgraph with names of your friends or f-o-f on the nodes. (Youcan use any plotting package or submit a handwritten graph.)
2. How many triangles are you already part of?
3. How many incomplete triangles are you part of? These are exactly the people who are friends of your friends but not (yet) yours aka ’potential’ friends 🙂
Note: You have to name people in the constructed graph. Feel free to use any method to get this answer: Face-Face meetings, Phone calls, Social Media, Zoom/Teams chats.
Solution:
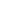



Reviews
There are no reviews yet.